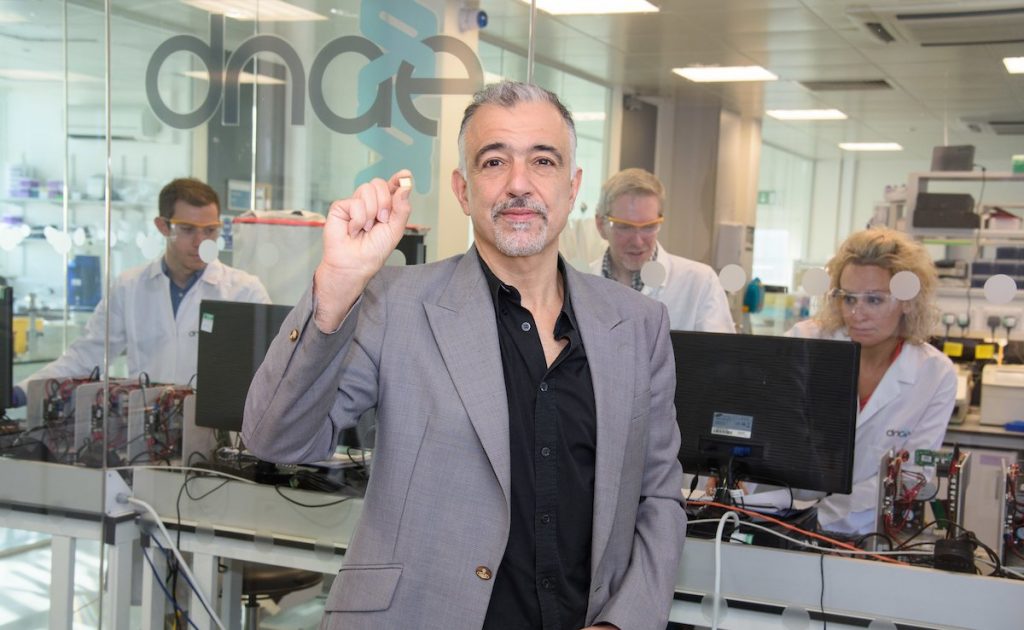
PISCES (pathogen identification from specimen, via capture extraction and sequencing) involves development and validation of DNAe’s test platform, branded Genalysis, and support for applications to the US Food and Drug Administration for marketing clearance.
“Genalysis platform will combine the ability to sequence the DNA of the infectious organism, in a sealed microchip based system, direct from clinical specimen, with analysis that enables actionable identification of the disease agent within a few hours,” said Imperial. “In influenza testing, the platform has the potential to ultimately identify the strain sub-type and anti-viral resistance markers.”
DNAe’s first intended product is a ‘blood-to-result’ diagnostic system for bloodstream infections leading to sepsis – set for commercial launch in 2018.
In 2014, BARDA awarded a $21.5m to DNAe’s US operation (then called nanoMR) to develop an automated sample preparation system that extracts and concentrates pathogens, such as bacteria, fungi and anthrax from infected blood samples.
DNAe acquired nanoMR in January 2015 to combine that pathogen capture system with its own DNA sequencing.
“The [combined] platform can be operated by users who are not specially-trained, enabling it to be used in near-to-patient clinical environments where sequencing has not been possible before,” said Sam Reed, president of DNAe’s US operation, “Unlike existing sequencing devices, the platform operates push-button directly from raw specimens such as blood or swabs.”
How it works
The basic technology looks for known stretches of DNA.
Its electronic sensor is a field-effect transistor whose gate is operated by hydrogen ions (protons) – such ions are released when a strand of DNA is extended by any of the four DNA nucleaotides A, G, C or T.
A key-like half-strand of DNA is created, called a primer, that will latch onto the DNA sequence being sought if it meets it.
This primer is put in with the sample, along with free A, G, C and T nucleaotides.
If the primer latches onto the sought sequence (called hybrisisation), free nucleaotides are encouraged to latch alongside to continue the match, releasing more ions, that the FET can detect, According to the firm. For dilute samples, the detection effect is enhanced if a DNA amplification technique is included in detection cell, and timing the output signals allows the original DNA concentration to be determined.
Different primers can be used on different parts of the DNA chip, and any heating required by the chemical reactions is provided by the chip.
BARDA is part of Assistant Secretary for Preparedness and Response (ASPR) in the US Department of Health and Human Services (HHS)
DNAe was founded by Professor Chris Toumazou, who invented the semiconductor DNA sequencing intellectual property behind the firm. It has operations in London, California and Washington DC, and its sequencing technology is licensed to Thermo-Fisher for research use.
 Electronics Weekly
Electronics Weekly